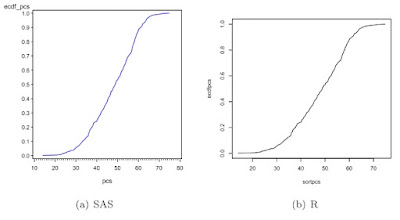
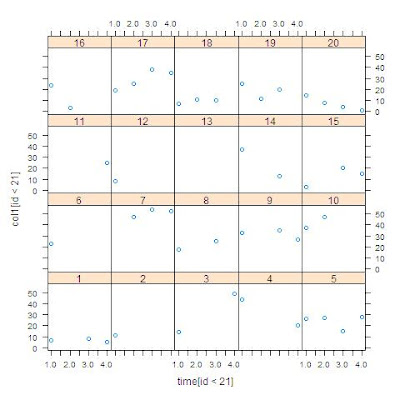
A related quantity is the Nelson-Aalen estimate of cumulative hazard. In addition to summarizing the hazard incurred by a particular timepoint, this quantity has been
used in missing data models (see White and Royston, 2009).
In addition to the usual SAS and R approaches to this, we also show Stata code.
SAS
It's very straightforward to calculate the Nelson-Aalen estimate in SAS. We assume the data is in the test data set generated in example 7.38.
ods output productlimitestimates=naout;
proc lifetest data=test nelson;
time time*event(0);
run;
options formchar="|----|+|---+=|-/\<>*";
options ls=64;
proc print data=naout; var time cumhaz; run;
In the forgoing, the formchar option makes SAS print in a format that can easily be cut-and-pasted into other software, such as Blogger's text entry window, as described here.
Obs time CumHaz
1 0.0000 0
2 0.5000 .
3 1.0000 .
4 1.0000 0.1111
5 2.0000 0.1736
6 2.0000 .
7 3.0000 0.2450
8 4.0000 0.3220
9 5.0000 .
10 6.0000 0.4129
11 7.0000 .
12 8.0000 0.5240
13 9.0000 0.6490
14 10.0000 .
15 12.0000 0.8156
16 14.0000 1.0156
17 14.0000 .
18 17.0000 .
19 20.0000 1.5156
20 21.0000 .
Note that SAS shows the Nelson-Aalen estimator once per time point, and only when there is a failure at that time. We may need the estimated value for all observed time points in our data, as well as to use the estimate with our original data. This requires some additional work. First, we sort our data and the new naout data set by time, removing rows from the above printout where the cumulative hazard is 0, using the where data set option (section A.6.3). Then we merge the two data sets.
proc sort data=test; by time; run;
proc sort data=naout (where=(cumhaz ne .)); by time; run;
data t2;
merge test (in=keepme) naout (keep = time cumhaz);
* the keep option drops unneeded variables;
by time;
if keepme;
* deletes any extra rows from the naout dataset;
retain cumhazlast 0;
* remember most recent value of this;
if cumhaz eq . then cumhaz = cumhazlast;
* plug in most recent value of cumhazlast,
if cumhaz is missing;
cumhazlast = cumhaz;
* the current value of cumhaz is now the most recent;
run;
proc print data=t2;
var time event cumhaz;
run;
This generates the desired result:
Obs time event CumHaz
1 0.5000 0 0
2 1.0000 1 0.1111
3 1.0000 1 0.1111
4 2.0000 1 0.1736
5 2.0000 0 0.1736
6 3.0000 1 0.2450
7 4.0000 1 0.3220
8 5.0000 0 0.3220
9 6.0000 1 0.4129
10 7.0000 0 0.4129
11 8.0000 1 0.5240
12 9.0000 1 0.6490
13 10.0000 0 0.6490
14 12.0000 1 0.8156
15 14.0000 0 1.0156
16 14.0000 1 1.0156
17 17.0000 0 1.0156
18 20.0000 1 1.5156
19 21.0000 0 1.5156
R
While the survfit() command can be used to create a table of all of the survival function and Nelson-Aalen estimates at each event point, it is slightly harder in R to associate these with the original data. Here we craft a function in R that does this housekeeping, using the fact that the Nelson-Aalen is just the negative log of the survival function (after specifying the type="aalen" option.
calcna = function(time, event) {
na.fit = survfit(coxph(Surv(time,event)~1), type="aalen")
jumps = c(0, na.fit$time, max(time))
# need to be careful at the beginning and end
surv = c(1, na.fit$surv, na.fit$surv[length(na.fit$surv)])
# apply appropriate transformation
neglogsurv = -log(surv)
# create placeholder of correct length
naest = numeric(length(time))
for (i in 2:length(jumps)) {
naest[which(time>=jumps[i-1] & time<=jumps[i])] =
neglogsurv[i-1] # snag the appropriate value
}
return(naest)
}
This can be be used as follows, where we use the time and event objects created in example 7.38.
newna = calcna(time, event)
cbind(time, newna)
time newna
[1,] 0.5 0.0000000
[2,] 1.0 0.1111111
[3,] 1.0 0.1111111
[4,] 2.0 0.1736111
[5,] 2.0 0.1736111
[6,] 3.0 0.2450397
[7,] 4.0 0.3219628
[8,] 5.0 0.3219628
[9,] 6.0 0.4128719
[10,] 7.0 0.4128719
[11,] 8.0 0.5239830
[12,] 9.0 0.6489830
[13,] 10.0 0.6489830
[14,] 12.0 0.8156496
[15,] 14.0 1.0156496
[16,] 14.0 1.0156496
[17,] 17.0 1.0156496
[18,] 20.0 1.5156496
[19,] 21.0 1.5156496
We also describe how to calculate the Nelson-Aalen estimate in Stata. First we create a dataset in the appropriate format. The foreign library in R (sections 1.1.5, 1.2.2) can do this directly. In SAS we would need to export to a format that Stata can read.
library(foreign)
write.dta(data.frame(time=time, event=event), "forstata.dta")
Stata
We can then read in the dataset from R:
. use forstata
(Written by R. )
. stset time, failure(event)
failure event: event != 0 & event < .
obs. time interval: (0, time]
exit on or before: failure
------------------------------------------------------------------------------
19 total obs.
0 exclusions
------------------------------------------------------------------------------
19 obs. remaining, representing
11 failures in single record/single failure data
156.5 total analysis time at risk, at risk from t = 0
earliest observed entry t = 0
last observed exit t = 21
. sts list, cumhaz
failure _d: event
analysis time _t: time
Beg. Net Nelson-Aalen Std.
Time Total Fail Lost Cum. Haz. Error [95% Conf. Int.]
-------------------------------------------------------------------------------
.5 19 0 1 0.0000 0.0000 . .
1 18 2 0 0.1111 0.0786 0.0278 0.4443
2 16 1 1 0.1736 0.1004 0.0559 0.5393
3 14 1 0 0.2450 0.1232 0.0915 0.6565
4 13 1 0 0.3220 0.1453 0.1330 0.7795
5 12 0 1 0.3220 0.1453 0.1330 0.7795
6 11 1 0 0.4129 0.1714 0.1830 0.9313
7 10 0 1 0.4129 0.1714 0.1830 0.9313
8 9 1 0 0.5240 0.2042 0.2441 1.1248
9 8 1 0 0.6490 0.2394 0.3149 1.3375
10 7 0 1 0.6490 0.2394 0.3149 1.3375
12 6 1 0 0.8156 0.2917 0.4046 1.6442
14 5 1 1 1.0156 0.3537 0.5132 2.0099
17 3 0 1 1.0156 0.3537 0.5132 2.0099
20 2 1 0 1.5156 0.6125 0.6865 3.3463
21 1 0 1 1.5156 0.6125 0.6865 3.3463
-------------------------------------------------------------------------------