In honor of
World Statistics Day and the
read paper that my co-authors Chris Wild, Maxine Pfannkuch, Matt Regan, and I are presenting at the Royal Statistical Society today, we present the R code to generate a combination dotplot/boxplot that is useful for students first learning statistics. One of the over-riding themes of the paper is that introductory students (be they in upper elementary or early university) should keep their eyes on the data.
When describing distributions, students often are drawn to the most visually obvious aspects: median, quartiles and extremes. These are the ingredients of the basic boxplot, which is often introduced as a graphical display in introductory courses. Students are taught to calculate the quartiles, and this becomes one of the components of a boxplot.
One limitation of the boxplot is that it loses the individual data points. Given a dotplot (see
here for an example of one in Fathom), it is very easy to guesstimate and draw a boxplot by hand. Wild et al. argue that doing so is probably the best way of gaining an appreciation of just what a boxplot actually is. The boxplot provides a natural bridge between operating entirely in terms of what is seen in graphics to reasoning using summary statistics. Retaining the dots in the combination dotplot/boxplot provides a reminder that the box plot is just summarizing the raw data, thus preserving a connection to more concrete foundations.
Certainly, such a plot has a number of limitations. It breaks down when there are a large number of observations, and has redundancy not suitable for publication. But as a way to motivate informal inference, it has potential value. It's particularly useful when combined in an animation using multiple samples from a known population, as demonstrated
here (scroll through the file quickly).
In addition, the code, drafted by Chris and his colleagues Steve Taylor and Dineika Chandrananda at the
University of Auckland, demonstrates a number of useful and interesting techniques.
RBecause of the length of the code, we'll focus just on the
boxpoints3() function shown below. The remaining support code must be run first (and can be found at
http://www.math.smith.edu/sasr/examples/wild-helper.R). The original
boxpoints() function has a large number of options and configuration settings, which the function below sets at the default values.
# Create a plot from two variables, one continuous and one categorical.
# For each level of the categorical variable (grps) a stacked dot plot
# and a boxplot summary are created. Derived from code written by
# Christopher Wild, Dineika Chandrananda and Steve Taylor
#
boxpoints3 = function(x,grps,varnames1,varnames2,labeltext)
{
observed = (1:length(x))[!is.na(x) & !is.na(grps)]
x = x[observed]
grps = as.factor(grps[observed])
ngrps = length(levels(grps))
# begin section to align titles and labels
xlims = range(x) + c(-0.2,0.1)*(max(x)-min(x))
top = 1.1
bottom = -0.2
plot(xlims, c(bottom,top), type="n", xlab="", ylab="", axes=F)
yvals = ((1:ngrps)-0.7)/ngrps
text(mean(xlims), top, paste(varnames1, "by", varnames2, sep=" "),
cex=1, font=2)
text(xlims[1],top-0.05,varnames2, cex=1, adj=0)
addaxis(xlims, ylev=0, tickheight=0.03, textdispl=0.07, nticks=5,
axiscex=1)
text(mean(xlims), -0.2, varnames1, adj=0.5, cex=1)
# end section for titles and labels
for (i in 1:ngrps) {
xi = x[grps==levels(grps)[i]]
text(xlims[1], yvals[i]+0.2/ngrps,
substr(as.character(levels(grps)[i]), 1, 12), adj=0, cex=1)
prettyrange = range(pretty(xi))
if (min(diff(sort(unique(xi)))) >= diff(prettyrange)/75)
xbins = xi # They are sufficiently well spaced.
else {
xbins = round(75 * (xi-prettyrange[1])/diff(prettyrange))
}
addpoints(xi, yval=yvals[i], vmax=0.62/ngrps, vadd=0.075/ngrps,
xbin=xbins, ptcex=1, ptcol='grey50', ptlwd=2)
bxplt = fivenum(xi)
if(length(xi) != 0)
addbox(x5=bxplt, yval=yvals[i], hbxwdth=0.2/ngrps,
boxcol="black", medcol="black", whiskercol="black",
boxfill=NA, boxlwidth=2)
}
}
Most of the function tends to issues of housekeeping, in particular aligning titles and labels. Once this is done, the support functions
addpoints() and
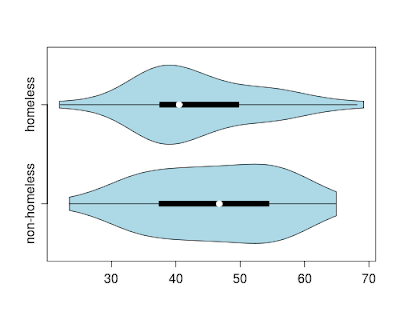
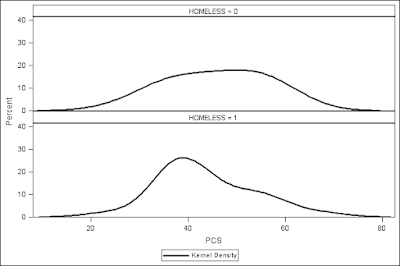
addbox() functions are called with the appropriate arguments. We can call the function using data from female subjects at baseline of the HELP study, comparing PCS (physical component scores) for homeless and non-homeless subjects.
source("http://www.math.smith.edu/sasr/examples/wild-helper.R")
ds = read.csv("http://www.math.smith.edu/r/data/help.csv")
female = subset(ds, female==1)
with(female,boxpoints3(pcs, homeless, "PCS", "Homeless"))
We see that the homeless subjects have lower PCS scores than the non-homeless (though the homeless group also has the highest score of either group in this sample).